Researchers Developed a Deep-Coverage, High-Throughput Method for Single-Cell Metabolomics
Date:2026-02-09
Single-cell metabolomics offers a powerful approach for characterizing metabolic heterogeneity at the individual-cell level that is obscured in bulk analyses. Mass spectrometry-based single-cell metabolomics enables the label-free, high-throughput detection of metabolites; however, current methods are not sensitive enough to detect low-abundance metabolites, and they show high technical variability and restricted coverage and annotation confidence.
In a study published in Nature Methods, research teams led by Prof. ZHU Zheng-Jiang at Shanghai Institute of Organic Chemistry (SIOC) of the Chinese Academy of Sciences develop an ion mobility-resolved mass cytometry technology for single-cell metabolomics (https://doi.org/10.1038/s41592-025-02970-2). By integrating novel data acquisition with advanced processing strategies, this single-cell metabolomics method achieves high throughput while maintaining deep coverage.
Researchers first established an ion mobility-resolved mass cytometry platform by integrating flow cytometry with ion mobility–mass spectrometry (IM–MS). The system continuously injects live cells into the IM–MS for direct electrospray ionization, transfers the pulsed single-cell ions that are generated into an ion trap for accumulation, and performs ion mobility separation with multidimensional measurements. To improve the detection of low-mass metabolites and reduce interference from cellular lipids, researchers implemented selective ion accumulation in the low-mass range on trapped ion mobility spectrometry. With this strategy, the abundance of low-mass ions was enhanced, increasing the sensitivity of detection for low-mass metabolites by 20-fold compared to the default condition. In addition, to address the low robustness of low-mass metabolites, a cell superposition strategy was developed. This strategy leverages distinctive ion signatures characterized by mass-to-charge ratio (m/z) and ion mobility values to aggregate identical ions across multiple cells. It enhanced peak detection and guided targeted ion extraction in individual cells, thereby improving robustness while maintaining single-cell resolution. This strategy elevated the median signal-to-noise ratio by 33-fold in peak detection and reduced metabolite dropout rates from 82% to 6% in quantification. Together, the selective ion accumulation and cell superposition enabled attomole-level sensitivity and a broad dynamic range at the single-cell level.
Then, researcher developed MetCell, an end-to-end computational tool optimized for ion mobility-resolved single-cell metabolomics. Overall, the technology detected over 5,000 metabolic peaks and annotated approximately 800 metabolites per cell, representing a 3–10-fold improvement over existing approaches. Notably, 389 metabolites were identified with level 1 confidence, which is the highest number reported at this level in single-cell metabolomics currently reported. Finally, using this technology, researchers curated a metabolic single-cell atlas containing 45,603 primary liver cells from aging mice. They demonstrated accurate cell type and subtype annotation using single-cell metabolomics and unveiled distinct metabolic states and heterogeneity of hepatocytes during aging.
This work was supported by National Natural Science Foundation of China, the Chinese Academy of Sciences, the Shanghai Municipal Science and Technology Commission, and the Shanghai Academy of Natural Sciences, among other funding sources.
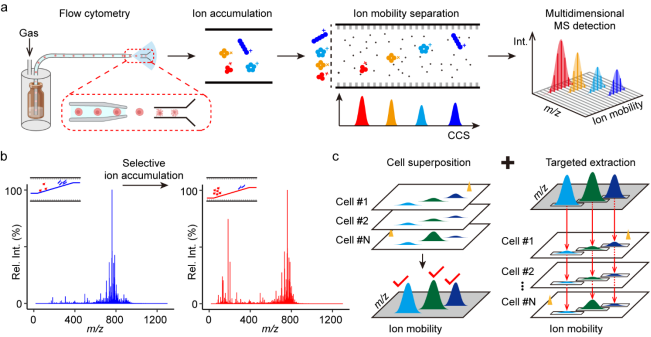
Figure 1. Ion mobility-resolved mass cytometry for single-cell metabolomics.
附件下载:
